Dunn Colin E. Biogeochemistry in Mineral Exploration
Подождите немного. Документ загружается.

individual species, groups or domains such as bacteria, archaea or eukaryotes in a soil
sample (Amann et al., 1995; Barns and Nierzwicki-Bauer, 1997).
Recently, an accurate quantitative analysis has been developed of the target DNA
from community DNA and RNA (so-called real-time PCR). This technique allows
quantifying how many copies of a particular gene are present in the soil sample
(Widada et al., 2002). These results can be correlated to the total amount of organ-
isms that express this gene in a sample and the rates of biogeochemical processes in
the soil samples. This makes real-time PCR a very powerful method to establish a
mechanistic link between the genetic composition of a microbial community and the
microbially mediated geochemical processes in soil materials that are relevant to
exploration geochemistry (Widada et al., 2002).
Assessing genetic diversity
To assess genetic diversity and identify key-species of microbial communities in
environmental samples, DNA libraries of unknown specie s are created (Fig. 12-2;
Barns and Nierzwicki-Bauer, 1997; Giraffa and Neviani, 2001). The DNA sequences
in these libraries can then be compared to other sequences deposited in global
sequence databases, which are an increa singly accurate and fast way of identification,
similar to the way that human DNA samples can be used to identify individuals in
criminal forensics.
Genetic fingerprinting
A number of techniques have been developed to obtain genetic fingerprints of
microbial communities in soil samples, and to relate these structures to environmental
conditions of different samples or treatments. The most commonly used techniques,
shown in Fig. 12-2, are as follows:
DGGE – denaturing gradient gel electrophoresis
TGGE – thermal gradient gel electrophoresis
SSCP – single strand conformation polymorphism
T-RFLP – terminal restriction fragment length polymorphism.
Initially, these methods were used to assess the phylogenetic relationships and
genetic diversity of microbial communities based on 16S rDNA or 18S rDNA, but
now they are also used to assess microbial communities based on functional genes
(Giraffa and Neviani, 2001; Widada et al., 2002).
1.
DGGE and TGGE are methods by which similar sized fragments of PCR-
amplified DNA can be resolved electrophoretically by their nucleotide composition
(Giraffa and Neviani, 2001; Widada et al., 2002). The double-stranded DNA
402
Exploration Geomicrobiology – the New Frontier
molecules have different melting behaviour and will stop at different pos itions
along the gel, as shown in Fig. 12-5. The emerging banding pattern is stained
to allow visualization and then analysed for similarity using cluster analysis
(Fig. 12-6) or non-metric multidimensional scaling (nMDS, Powell et al., 2003).
Thus, PCR-DGGE and -TGGE methods provide an ecological insight into the
structure of microbial communities in environmental samples, and may also be
used to identify key organisms (see Case Study 1) whi ch can be cloned, sequenced
and subsequently identified.
2.
SSCP also detects sequ ence variations between different amplified target DNA
fragments.
3.
T-RLFP is a method to study mixed populations in soil samples that is based on
restriction enzyme digestion of fluorescently marked PCR products.
4.
Phospholipid fatty acid (PLFA) analysis is an additional fingerprinting technique,
based on using cell constituents other than DNA or RNA ( Barns and Nierzwicki-
Bauer, 1997). PLFA are fatty acids present in the lipid bi-layer membranes of
living micro-organisms that are unique for specific groups of micro-organi sms.
Thus, PLFA analyses can be used to assess the diversity of a microbial community
and to evaluate changes in microbial community structures as a result of changing
biogeochemical conditions.
CASE STUDIES
Three case studies are presented to demo nstrate how classical and molecular tools
enhance the understanding of micro-organisms and microbially mediated processes
associated with the turnover of trace metals in soils and deeper soil materials, and
how this increased understanding may lead to the following:
(1)
The prediction of mineral transport and transformation pathways and the
location of secondary mineralized zones in the soil.
(2)
The development of biosensors for masked mineralization.
Case Study 1 – The geomicrobiological cycling of gold
This study assessed the ability of natural soil microbiota to mediate the
solubilization, transport and precipitation of Au in the Australian regolith, and is
taken from Reith et al. (2005), Reith and McPhail (2006), Reith et al. (2006) and
Reith and McPhail (2007).
Whereas it is now well established that micro-organisms play a key in role in the
cycling of major and trace elements in the environment, current evidence for the role
of micro-organisms in the biogeochemical cycling of Au is at best equivocal (Mossman
et al., 1999). Laboratory experiments using pure cultures of common bacteria such as
Bacillus subtilis or Bacillus megaterium have shown that these organisms can solubilize
403
Biogeochemistry in Mineral Exploration
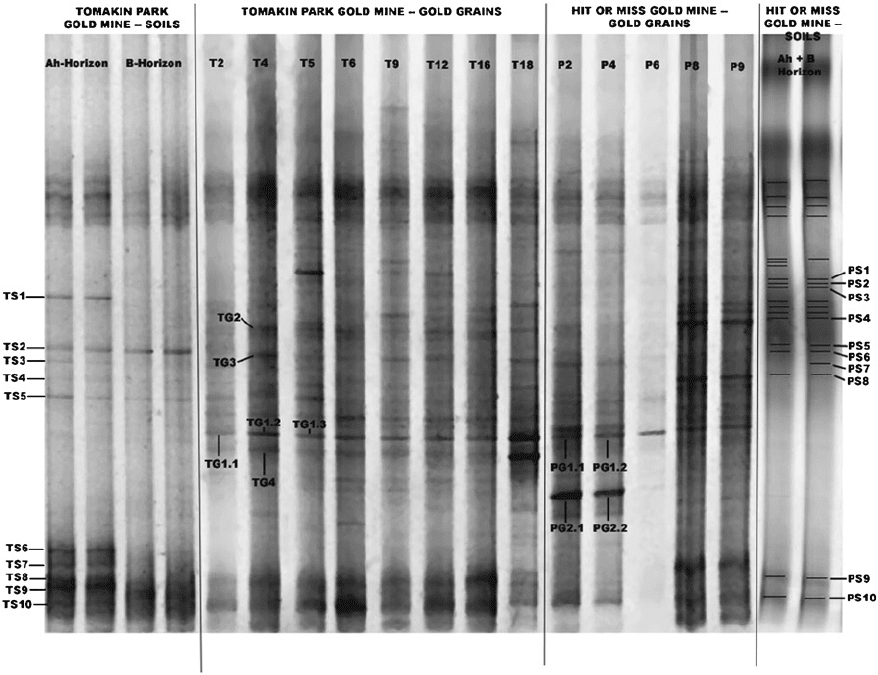
Fig. 12-5. DGGE (denaturing gradient gel electrophoresis) patterns obtained after amplifying the V3 region of the 16S ribosomal DNA
extracted from Au grains and surrounding soils collected at the Tomakin Park- (T) and the Hit or Miss (P) gold mines. Bands designated
with acronyms were excised from the gels, re-amplified and sequenced (Reith et al., 2006).
404 Exploration Geomicrobiology – the New Frontier
Au (Korobushkina et al., 1983). The ability of some bacteria (Escherichia coli,
Pseudomonas aeruginosa)andfungi(Aspergillus niger, Penicillium chrysogenum)to
precipitate mobile Au complexes and colloids in vitro has also been demonstrated
(Karthikeyan and Beveridge, 2002; Nakajima, 2003). Furthermore, morphological
evidence for a microbially mediated formation of Au grains and nuggets in soils and
deeper soil materials has been presented (Mann, 1992; Keeling, 1993; Bischoff, 1994,
1997). Using biochemical tools, Levchenko et al. (2002) demonstrated that a Au (I/III)-
redox couple acts as a metal centre in the membrane-bound enzyme NADH-oxidase
utilized by the common bacterium Micrococcus luteus during the oxidation of methane.
Whereas this limited evidence suggested that microbial biogeochemical mechanisms
may exist for the solubilization and precipitation of Au, the relative importance of
these mechanisms in the regolith compared to abiotic processes and the organisms
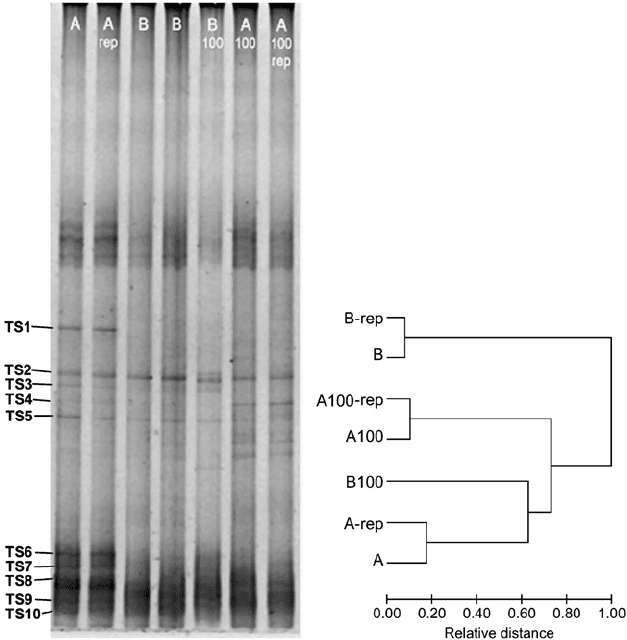
Fig. 12-6. Molecular fingerprinting analyses showing DGGE-patterns of 16S rDNA and
dendrogram of subsequent cluster analysis from auriferous (A, B) and non-auriferous (A100,
B100) soils from the Tomakin Park Gold Mine.
405Biogeochemistry in Mineral Exploration
driving these processes are not known. Thus, the aim of the research was to obtain
evidence for the presence and significance of a biologically mediated cycle of Au in the
regolith, and subsequently identify and quantify microbial processes that are important
in mediating the dispersion and concentration of Au.
Figure 12-7 shows, from the sequence of observations, that microbiota resident in
auriferous Australian soils and deeper regolith materials are capable of mediating a
geomicrobiological cycle of Au. The indigenous microbiota in biologically active soil
microcosms were able to solubilize up to 80 wt.% of the Au contained in these
materials during the first 50 days of incubation, after which the solubilized Au was
re-adsorbed by mineral- and organic soil fractions. In contrast, no Au was solubilized
in sterilized microcosms incubated under otherwise identical conditions. Molecular
(PCR-DGGE, cloning and sequencing of 16s rDNA) and physiological profiling
(CLPP) of bacterial communities during the incubation of the microcosms combined
with amino acids analyses indicated that changes in the structure of the bacterial
community from carbohydrate- to amino acid-utilizing populations occurred concur-
rently with, and appear to be linked to, the observed solubilization and re-precipitation
of Au.
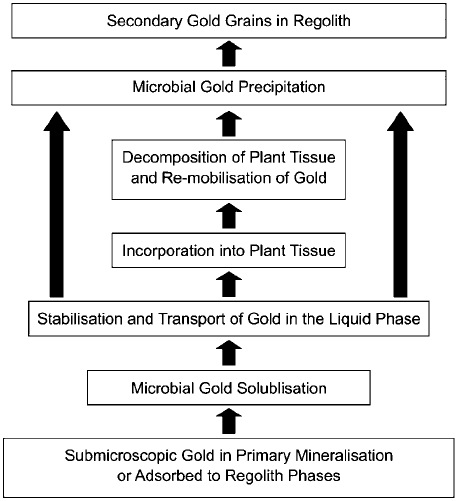
Fig. 12-7. Conceptual model linking the processes of Au solubilization, transport, precip-
itation and authigenic Au nugget formation to form a geo(micro)biological cycle for the
behaviour of Au in the regolith.
406 Exploration Geomicrobiology – the New Frontier
These results suggest the following model of Au solubilization and re-precipitation
in the soil microcosms (Fig. 12-8).
The bacterial community in the early stages of incubation apparently produced
surplus amino acids, which are known to solubilize native Au a nd stabilize it in
solution.
The bacterial community in the later stages of incubation utilized these ligands,
and as a result the Au complexes were destabilized and Au was re-precipitated to
the solid soil fractions.
Molecular profiling also allowed the differentiation of bacterial communities
from auriferous and adjacent non-auriferous soils (Fig. 12-6). These results in com-
bination with results of Bacillus cereus spore counts (which were up to 1000 times
higher in soils that displayed Au concentrations of 150–1000 ppb compared to soils
with background Au concentrations), and microcosms amended with dissolved
AuCl
4
–
, suggest that the presence of highly mobile Au in the regolith as observed
at many Australian sites may influence the composi tion of the resident microbiota.
The results indicate that a geomicrobial exploration technique may be developed,
where B. cereus spore counts are measured and used as a pre-screening method to
target areas useful for further sampling and complete geochemical analysis (Reith
et al., 2005).
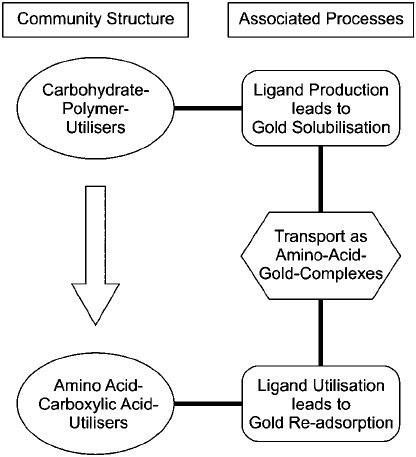
Fig. 12-8. Conceptual model linking Au solubilization and re-precipitation in microcosm
experiments with auriferous soils to the results of microbial community structure analyses.
407Biogeochemistry in Mineral Exploration

Scanning electron microscopy (SEM) revealed bacterial pseudomorphs on
untreated secondary Au grains from two field sites used in this study (Fig. 12-9).
The presence of active bacterial biofilms on the surface of Au grains was confirmed
using confocal stereo laser microscopy (CSLM) combined with nucleic acid staining.
Molecular profiling showed that unique, site-specific bacterial communities are
associated with these Au grains, which differed from those dominating the surrounding
soils (Figs. 12.4 and 12.5). 16S ribosomal DNA clones belonging to the genus Ralstonia
and bearing 99% similarity to Ralstonia metallidurans were present on all 16S rDNA
positive Au grains from both locations, but were not detected in the surrounding soils
(Fig. 12-5). The ability of R. metallidurans to actively accumulate Au from solution
was then successfully tested suggesting that R. metallidurans may contribute to the
formation of secondary Au grains and nuggets in the regolith.
These studi es have shown for the first time that microbiota resident in Australian
regolith are capable of actively mediating a biogeochemical cycle of Au in the
environment, and that the micro bially induced turnover of Au in the regolith may be
rapid. Furthermore, there is evidence for a number of processes and organisms that
may be associated with this cycle. Future experiments will use molecular microarrays
to assess Au solubilizing and precipitating organisms, such as R. metallidurans,in
order to understand the specific biochemical mechanisms regulating Au solubilizat-
ion and precipitation. This may enable the development of specific gene probes for
genes associated with the biogeochemical cycle of Au. Novel biosen sor technology
(as shown in Case Study 3) in combination with molecular characterization of
Au-transforming organisms and associated biochemical processes will then have
the potential to facilitate the development of molecular biosensors specific to Au for
use in the field.
Fig. 12-9. Secondary electron micrographs of surface features of secondary Au grains from
the Hit or Miss Gold Mine in the Palmer River Goldfields in north eastern Queensland,
Australia. (a) ‘Bacterioform’ Au consisting of bacterial cell-like structures on the surface of the
grain (scale bar ¼ 5 mm) and (b) Initial stages of ‘bacterioform’ Au formation in a surface
depression of a Au grain (scale bar ¼ 20 mm).
408 Exploration Geomicrobiology – the New Frontier
Case Study 2 – Microbial diversity in soils and sediments contaminated with Zn
and Cu
Brim et al. (1999) examined bacterial communities of Zn-contaminated soils
from Belgium using a combination of molecular and classical methods. From an
exploration geomicrobiology perspective these results are interesting, because they
show that the composition of the resident bacterial community was dominantly
influenced by the concentration and speciation of Zn in these soils, as may be the
case in soils overlying buried mineralization. The site had been monitored for two
decades, and previous studies using cultivation-based techniques had shown that
Ralstonia eutropha-like strains had dominated in the microbial community in the
soil, i.e., in previous studies up to 40% of cultured cells belonged to the Ralstonia
group.
In this study using a colony-forming-units technique, 10
4
–10
5
g
–1
culturable soil
bacteria were counted in a number similar to results of previous studies. However, 23
of the isolates belonged to the Arthrobacter group of bacteria, but no R. eutropha-like
isolates were detected. Most of the isolates were Zn tolerant but only seven were
considered Zn resistant. Sequences obtained from 16S rDNA clones from a clone
library, established from the soil community DNA, were affiliated with a number of
different groups such as a- and b-proteobacteria and the Cytophaga- Flexibacter-
Bacteroides group. Molecular profiling, using TGGE, of amplified 16S rDN A from
isolates, soil clones and the extracted soil community DNA showed that isolates and
clones only represented a pa rt of the microbial population of these soils. Arthrobacter
was the dominant band when 16S rDNA had been amplified from DNA extracted
from the soil in a procedure with a bead -beating step, but was absent or faint when
soil DNA was extracted without bead beating. From these results, Brim et al. (1999)
concluded that the microbial community in previous studies was highly resistant to
Zn toxicity concentration but was replaced with a less resistant bacterial microflora.
They suggest ed that Zn toxicity levels had been reduced because of the activity of
the Zn resistant R. eutropha-like bacteria that had led to the formation of insoluble
Zn carbonates. The results also showed that molecular techniques such as TGGE
in combination with classical techniques are powerful tools to assess microbial
community structures, as they allow monitoring of the temporal and spatial changes
of microbial communities and may facilitate the identification of key organisms, i.e.,
R. eutropha- and Arthrobacter-like strains.
In another study published by Konstantinidis et al. (2003) Ralstonia-and
Arthrobacter-like strains were the only Cu resistant isolates cultivated from lake
sediments contaminated with 200–5500 ppm Cu. T-RFLP showed that a numb er of
OTUs (operational taxonomic units), among them Ralstonia sp., were present uni-
versally the entire width of the sedim ent. A number of other studi es also demon-
strated the presence Ralstonia sp. and Arthrobacter sp. in samples with elevated
concentrations of heavy metals and may as such be tried as bio-indicator organisms
in geomicrobial mineral exploration.
409
Biogeochemistry in Mineral Exploration
Case Study 3 – Bacterial biosensors as alternatives for measuring heavy metals
in soil extracts
This study tested the applicability of metal-specific whol e-cell bacterial sensors for
the analysis of arsenite and Hg in soil extracts from polluted soils by comparing them
to resul ts from ICP-AES analyses. The luminescence-based bacterial sensor strain
Pseudomonas fluorescens OS8 (pTPT11) was used for Hg detection and Pseudomonas
fluorescens OS8 (pTPT31) for arsenite detection. Petanen and Romantschuk (2002)
spiked three different soil types (humus, mineral and clay) with 1100 or 500 mgg
–1
(ppm) of dissolved Hg(II) or As(III). They took samples after 1, 14 and 30 days
of incubation and extracted with water, ammonium acetate, hydrogen peroxide
and nitric acid to represent water soluble, bioavailable, organic matter-bound and
residual fractions, respectively.
Concentration results with chemical and biosensor analysis were similar in the
case of Hg-spiked samples. However, the lowest Hg concentration measured in the
soil extracts using the bacterial sensor was 0.003 mgkg
–1
(ppb) which was consid-
erably lower than by the chemical method 0.05 mgkg
–1
(ppb). The sensor strain with
pTPT31 for arsenite had a useful detection range similar to that of chemical methods.
Thus, they demonstrated that the bacterial sensors were sufficiently sensitive to
measure concentrations of As and Hg that are relevant to mineral exploration
in their soil extracts. Recently, other authors found similar low-detection limits in
biosensors for Cu, Co, Zn, Pb and Cd (Tibazarwa et al., 2001; Rensing and Maier,
2003).
FUTURE TECHNOLOGIES FOR BIO-PROSPECTING
There are several emerging molecular techniques that have the potential to
become the basis of future bio-prospecting technologies – these are nucleic acid
probes, molecular micro-arrays, biosensors and immuno-assays. These techniques
will provide the platform on which bio-prospecting test kits will be developed for the
analyses of soil samples in the field. It is expected that within the next ten years field
test kits will be developed that permit analyses of up to several hundred samples
per day. Use of these methods to supplement traditional chemi cal methods will have
a number of advantages for mineral exploration in that, apart from providing
an additional layer of information, they are rapid, inexpensive, simple to perform,
portable, highly sensitive and selective and thereby allow rapid assessment of
potential mineralization in the field. These techniques will not replace traditional
chemical analysis techniques but will provide an additional level of information and
they represent an additional tool in the mineral explorationist’s tool box. Molecular
techniques will add value to existing techniques, most importantly ‘green field’
exploration where they can narrow down the final drilling target in relatively cheap
and rapid ways.
410
Exploration Geomicrobiology – the New Frontier
The iden tification of bio-indicator organisms or genes present in elevated
numbers in soil samples with anomalous trace metals concentrations may lead to the
development of specific nuclei c acid probes for these bio-indicators that can be used
directly for bio-prospecting. Nucleic acid probes are complementary to signature
sequences of functional or ribosomal DNA or RNA of particular species or groups
of organisms and are fluorescently- or isotopically labelled allowing for their detec-
tion at very low concentration (Barns and Nierzwicki-Bauer, 1997). Nucleic acid
probes can bind to bulk community DNA or RNA bound in filter membranes
making it possibl e to quantify the amoun t and activity of particular bio-indicator
organisms (Barns and Nierzwi cki-Bauer, 1997).
Up to several hundred thousan d nucleic acid probe s can be used together as a
molecular micro-array, as shown in Fig. 12-10 (Widada et a l., 2002; Zhou and
Thomps on, 2002;, Zhou (2003) ; Bae and Park, in press). Molecular micro-arrays,
which all ow researchers t o study comple x microbial communities, comprise one
of the new tools in m olecular m icrobiology. Micro-arrays are powerful tools
for detection of multiple genes from soil samples, g iving a vast amount of i n-
formation relating to the phylogenetic and/or functional structure of microbial
ecosystems.
Expression micro-arrays are used to assess gene expression from microbial
cultures or environm ental samples. This type of micro-array provides information
relating to the activity of the microbe(s), i.e., ‘listening-in’ to the message provided by
the microbes. Furthermore, key genes associated with environmental attributes can
be identified. For example, genes can be identified that are important when a species
is subjected to elevated concentrations of heavy meta ls. Identification of genes
that facilitate heavy metal transformations may then lead to the development of
micro-arrays with specific probes for these genes. These can then be applied as
exploration tools after RNA extraction from soil samples.
Heavy metal specific bacterial sensors provide another promising bio-prospecting
tool, allowing the measurement of mobile heavy metals in soil samples to ppb levels
(Tibazarwa et al., 2001; Rensing and Maier, 2003). This technique, which is high-
lighted in Case Study 3, was initially developed for ecotoxicological testing of heavy
metal polluted sites in Europe and the US to distinguish between mobile and
bioavailable, and non-mobile and non-bioavailable fractions of heavy metals in soils
and sediments (Rensi ng and Maier, 2003). Metal specific bacterial sensors have been
developed for a number of metals and metalloids such as As, Cu, Ni, Co, Zn, Pb, Hg
and Cd, and could be tested as exploration tools in bio-prospecting (Rensing and
Maier, 2003). Such biosensors are able to distinguish between mobile transported
heavy metals and in-situ bac kground concentrations much better than by wet chem-
ical methods (Rensing and Maier, 2003). This is likely to be of specific interest to
geochemical exploration.
Antibody-based or immuno-assays offer an alternative approach for metal ion
detection in natural samples. An immuno-assay makes use of the binding between an
antigen and its homologous antibody in order to quantify the specific antigen or
411
Biogeochemistry in Mineral Exploration
